Sep 28, 2018 · newsletter
Snorkel: Rapid Training Data Creation with Weak Supervision
Machine learning (and more specifically, deep learning) techniques are recognized for their ability to obtain high accuracy for a variety of classification problems. Deep learning models are more complex than most traditional models and often have thousands of parameters and commensurately require more labeled training data. Obtaining this data can be an expensive and time-consuming process, and often requires domain experts to help with the labeling process. Many areas of machine learning that are motivated by this bottleneck could be broadly categorized into four areas: active learning, semi-supervised learning, weak supervision and transfer learning. Snorkel belongs to the weak supervision class and is effective for tasks like natural language processing. The basic idea is to allow users to provide supervision at a higher level than case-by-case labeling, and then to use various statistical techniques to deal with the noisier labels that are obtained. Working with domain experts, the Snorkel team has found that this type of supervision is easier and faster to provide. And surprisingly, by getting large volumes of lower-quality supervision in this way, one can train higher-quality models at a fraction of the time and cost.
Snorkel is a system built around the data programming paradigm for rapidly creating, modeling, and managing training data. It lets one use weak supervision sources like heuristics, external knowledge bases, crowdsourced labels, and more. These are called weak supervision sources because they may be limited in accuracy and coverage. All of these get combined in a principled manner to produce a set of probability-weighted labels. The end model is then trained on the generated labels.
Snorkel workflow
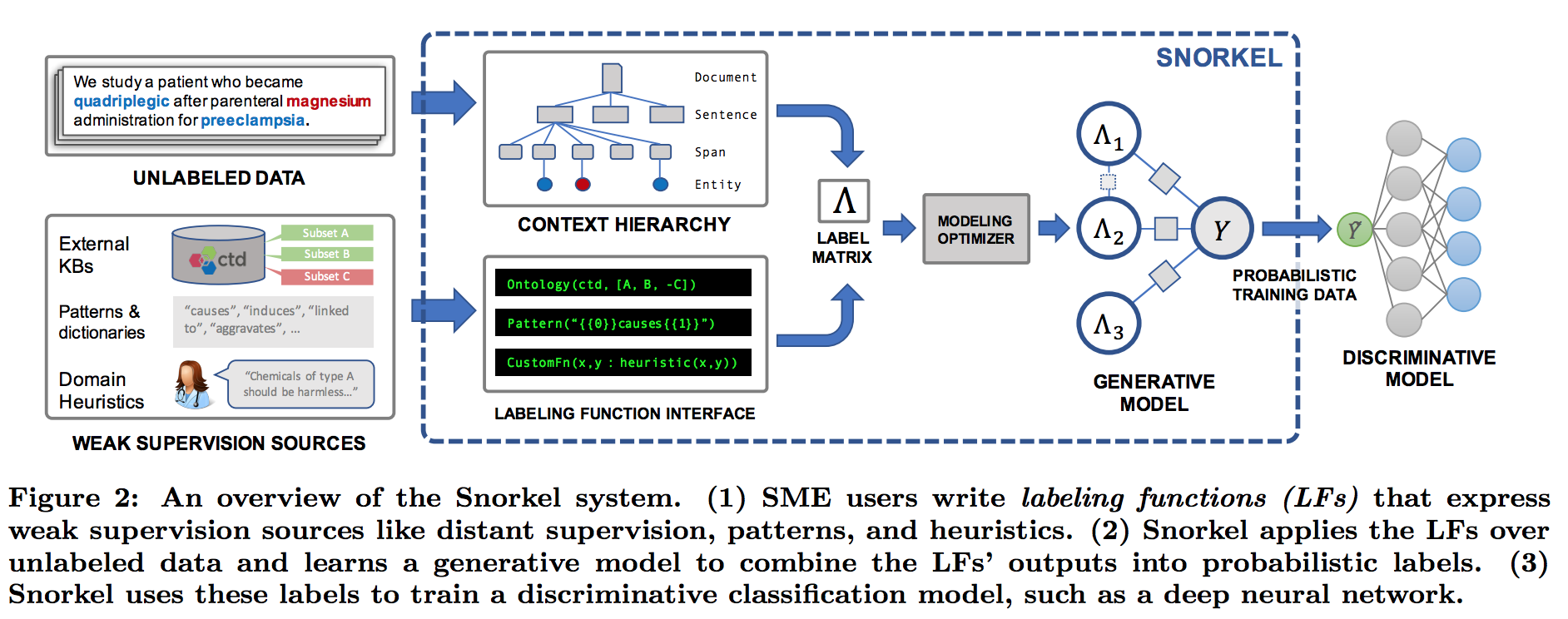
Snorkel’s workflow has three main stages:
- Writing Labeling Functions: Instead of hand-labeling large quantities of training data, users write labeling functions which capture patterns-based heuristics, connect with external knowledge bases (distant supervision), and so on. A labeling function is a Python method which, given an input, can either output a label or abstain. For instance, we can write a labeling function that checks if the word “causes” appears between the chemical and disease mentions. If it does, it outputs True if the chemical mention is first and False if the disease mention is first. If “causes” does not appear, it outputs None, indicating abstention. In addition, Snorkel includes a number of declarative labeling functions that can be used out of the box.
Example labeling function
 2. Modeling Accuracies and Correlations: Next, Snorkel automatically learns a generative model over the labeling functions, which allows it to estimate their accuracies and correlations. This step uses no ground-truth data, learning instead from the agreements and disagreements of the labeling functions.
2. Modeling Accuracies and Correlations: Next, Snorkel automatically learns a generative model over the labeling functions, which allows it to estimate their accuracies and correlations. This step uses no ground-truth data, learning instead from the agreements and disagreements of the labeling functions.
- Training a Discriminative Model: The output of Snorkel is a set of probabilistic labels that can be used to train a wide variety of state-of-the-art machine learning models, such as popular deep learning models. While the generative model is essentially a re-weighted combination of the user-provided labeling functions — which tend to be precise but low-coverage - discriminative models can retain this precision while learning to generalize beyond the labeling functions, increasing coverage and robustness on unseen data.
One thing to note is that it is quite possible for a user to write labeling functions that are statistically dependent (i.e., correlated). Modeling such dependencies is crucial because it affects the estimates of true labels. To help with this, Snorkel automatically selects which dependencies to model using an estimator that relies on a hyper-parameter - epsilon. With large values of epsilon, no correlations are included, and as we reduce the value progressively more correlations are added, starting with the strongest.
Snorkel is currently being used on text extraction applications for medical records at the Deparment of Veterans Affairs, to mine scientific literature for adverse drug reactions in collaboration with the Federal Drug Administration, and to comb through everything from surgical reports to after-action combat reports for valuable structured data. More details about the project can be found here.